fq2bam_meth
Generate BAM/CRAM output given one or more pairs of FASTQ files from bisulfite sequencing (BS-Seq). Can also optionally generate a BQSR report.
The tool fq2bam_meth is a fast, accurate algorithm for mapping methylated DNA sequence reads to a reference genome, performing local alignment, and producing alignment for different parts of the query sequence. It implements the baseline tool bwa-meth [1] [2] in a performant method using fq2bamfast (BWA-MEM + GATK) as a backend for processing on GPU.
fq2bam_meth is the Parabricks wrapper for bwa-meth, which will sort the output and can mark duplicates and recalibrate base quality scores in line with GATK best practices.
The Parabricks fq2bam_meth tool is capable of handling longer reads and is less sensitive to errors than other alignment algorithms. We enable fast and accurate whole-genome bisulfite sequencing (WGBS) to detect DNA-methylation at the single base pair level [3].
Some of the advantages of using fq2bam_meth over similar tools include:
It is faster than many other BS-Seq alignment algorithms, making it the ideal choice for high-throughput analysis.
It maintains compatibility with existing CPU-based tools.
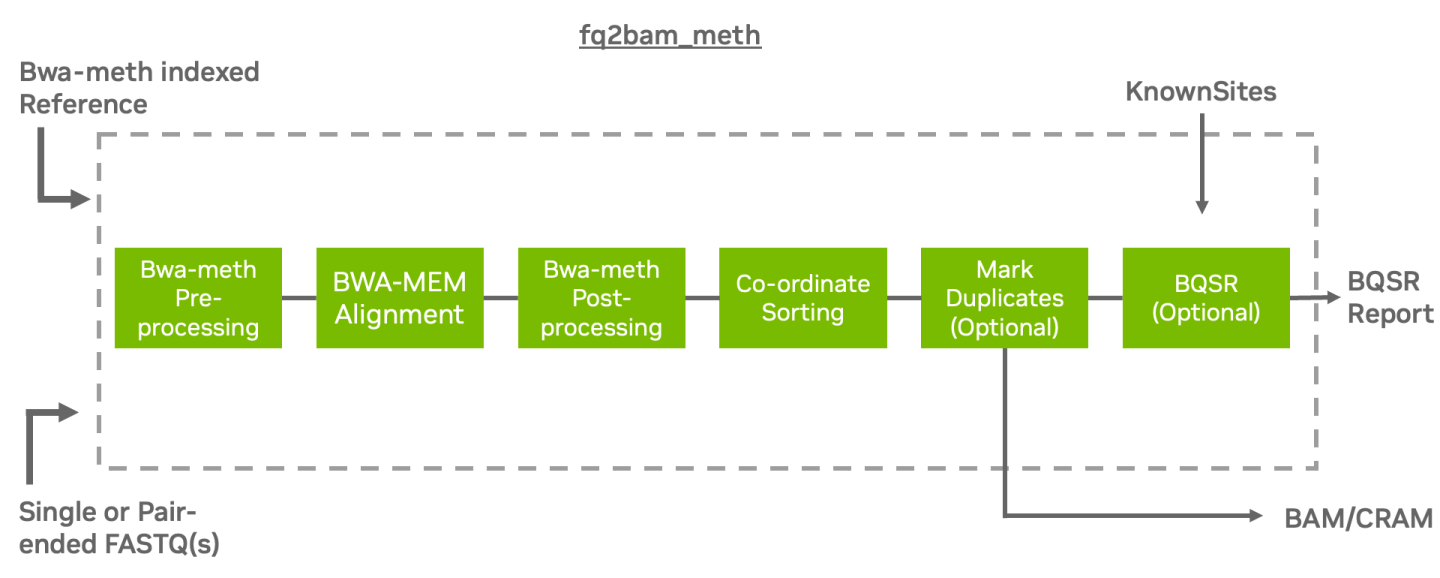
fq2bam_meth uses an accelerated version of BWA-MEM to generate BAM/CRAM output
given one or more pairs of FASTQ files from BS-Seq. The user can turn-off marking
of duplicates by adding the --no-markdups option. The BQSR step is only performed if the
--knownSites input and --out-recal-file output options are provided; doing so will also
generate a BQSR report.
Prior to running alignment, the reference genome must be converted using baseline bwa-meth.
The bwa-meth indexing step produces a reference fasta file with a name formatted as fasta.bwameth.c2t.
The indexing preparation step requires running bwameth.py index $REF.fasta.
Baseline bwa-meth requires baseline BWA-MEM to be in the user's path for indexing functionality.
Note that indexing is a time-consuming prerequisite that should only need to be completed once per reference genome.
The bwameth.py script can be found here.
# This command assumes all the inputs are in INPUT_DIR and all the outputs go to OUTPUT_DIR.
docker run --rm --gpus all --volume INPUT_DIR:/workdir --volume OUTPUT_DIR:/outputdir \
--workdir /workdir \
nvcr.io/nvidia/clara/clara-parabricks:4.3.1-1 \
pbrun fq2bam_meth \
--ref /workdir/${REFERENCE_FILE} \
--in-fq /workdir/${INPUT_FASTQ_1} /workdir/${INPUT_FASTQ_2} \
--knownSites /workdir/${KNOWN_SITES_FILE} \
--out-bam /outputdir/${OUTPUT_BAM} \
--out-recal-file /outputdir/${OUTPUT_RECAL_FILE}
The commands below are the bwa-meth-0.2.7, bwa-0.7.15, and GATK4 counterpart of the Parabricks command above. The output from these commands will be identical to the output from the above command. See the Output Comparison page for comparing the results.
Set --bwa-options="-K 10000000" in fq2bam_meth and -K 10000000 in baseline to produce compatible pair-ended results.
fq2bam_meth will not strip _R1 and _R2 from read names during preprocessing like baseline bwa-meth.
# Run bwa-meth and pipe the output to create a sorted BAM.
$ python bwa-meth.py \
--read-group '@RG\tID:sample_rg1\tLB:lib1\tPL:bar\tSM:sample\tPU:sample_rg1' \
--reference <INPUT_DIR>/${REFERENCE_FILE} <INPUT_DIR>/${INPUT_FASTQ_1} <INPUT_DIR>/${INPUT_FASTQ_2} \
-t 32 -K 10000000 | \
gatk SortSam \
--java-options -Xmx30g \
--MAX_RECORDS_IN_RAM 5000000 \
-I /dev/stdin \
-O cpu.bam \
--SORT_ORDER coordinate
# Mark duplicates.
$ gatk MarkDuplicates \
--java-options -Xmx30g \
-I cpu.bam \
-O mark_dups_cpu.bam \
-M metrics.txt
# Generate a BQSR report.
$ gatk BaseRecalibrator \
--java-options -Xmx30g \
--input mark_dups_cpu.bam \
--output <OUTPUT_DIR>/${OUTPUT_RECAL_FILE} \
--known-sites <INPUT_DIR>/${KNOWN_SITES_FILE} \
--reference <INPUT_DIR>/${REFERENCE_FILE}
Run GPU-accelerated bwa-meth compatible alignment, co-ordinate sorting, marking duplicates, and Base Quality Score Recalibration to convert bisulfite reads from FASTQ to BAM/CRAM.
Input/Output file options
- --ref REF
- --in-fq [IN_FQ ...]
- --in-se-fq [IN_SE_FQ ...]
- --in-fq-list IN_FQ_LIST
- --knownSites KNOWNSITES
- --interval-file INTERVAL_FILE
- --out-recal-file OUT_RECAL_FILE
- --out-bam OUT_BAM
- --out-duplicate-metrics OUT_DUPLICATE_METRICS
- --out-qc-metrics-dir OUT_QC_METRICS_DIR
Path to the reference file. We will automatically look for <filename>.bwameth.c2t. Converted fasta reference must exist from prior conversion with baseline bwa-meth (default: None)
Option is required.
Path to the pair-ended FASTQ files followed by optional read groups with quotes (Example: "@RG\tID:foo\tLB:lib1\tPL:bar\tSM:sample\tPU:foo"). The files must be in fastq or fastq.gz format. All sets of inputs should have a read group; otherwise, none should have a read group, and it will be automatically added by the pipeline. This option can be repeated multiple times. Example 1: --in-fq sampleX_1_1.fastq.gz sampleX_1_2.fastq.gz --in-fq sampleX_2_1.fastq.gz sampleX_2_2.fastq.gz. Example 2: --in-fq sampleX_1_1.fastq.gz sampleX_1_2.fastq.gz "@RG\tID:foo\tLB:lib1\tPL:bar\tSM:sample\tPU:unit1" --in-fq sampleX_2_1.fastq.gz sampleX_2_2.fastq.gz "@RG\tID:foo2\tLB:lib1\tPL:bar\tSM:sample\tPU:unit2". For the same sample, Read Groups should have the same sample name (SM) and a different ID and PU. (default: None)
Path to the single-ended FASTQ file followed by optional read group with quotes (Example: "@RG\tID:foo\tLB:lib1\tPL:bar\tSM:sample\tPU:foo"). The file must be in fastq or fastq.gz format. Either all sets of inputs have a read group, or none should have one, and it will be automatically added by the pipeline. This option can be repeated multiple times. Example 1: --in-se-fq sampleX_1.fastq.gz --in-se-fq sampleX_2.fastq.gz . Example 2: --in-se-fq sampleX_1.fastq.gz "@RG\tID:foo\tLB:lib1\tPL:bar\tSM:sample\tPU:unit1" --in-se-fq sampleX_2.fastq.gz "@RG\tID:foo2\tLB:lib1\tPL:bar\tSM:sample\tPU:unit2" . For the same sample, Read Groups should have the same sample name (SM) and a different ID and PU. (default: None)
Path to a file that contains the locations of pair-ended FASTQ files. Each line must contain the location of two FASTQ files followed by a read group, each separated by a space. Each set of files (and associated read group) must be on a separate line. Files must be in fastq/fastq.gz format. Line syntax: <fastq_1> <fastq_2> <read group> (default: None)
Path to a known indels file. The file must be in vcf.gz format. This option can be used multiple times. (default: None)
Path to an interval file in one of these formats: Picard-style (.interval_list or .picard), GATK-style (.list or .intervals), or BED file (.bed). This option can be used multiple times. (default: None)
Path of a report file after Base Quality Score Recalibration. (default: None)
Path of a BAM/CRAM file. (default: None)
Option is required.
Path of duplicate metrics file after Marking Duplicates. (default: None)
Path of the directory where QC metrics will be generated. (default: None)
Tool Options:
- --max-read-length MAX_READ_LENGTH
- --min-read-length MIN_READ_LENGTH
- -L INTERVAL, --interval INTERVAL
- --bwa-options BWA_OPTIONS
- --no-warnings
- --filter-flag FILTER_FLAG
- --skip-multiple-hits
- --align-only
- --no-markdups
- --fix-mate
- --markdups-assume-sortorder-queryname
- --markdups-picard-version-2182
- --monitor-usage
- --optical-duplicate-pixel-distance OPTICAL_DUPLICATE_PIXEL_DISTANCE
- --read-group-sm READ_GROUP_SM
- --read-group-lb READ_GROUP_LB
- --read-group-pl READ_GROUP_PL
- --read-group-id-prefix READ_GROUP_ID_PREFIX
- -ip INTERVAL_PADDING, --interval-padding INTERVAL_PADDING
- --standalone-bqsr
- --set-as-failed SET_AS_FAILED
- --do-not-penalize-chimeras
Maximum read length/size (i.e., sequence length) used for bwa and filtering FASTQ input (default: 480)
Minimum read length/size (i.e., sequence length) used for bwa and filtering FASTQ input (default: 10)
Interval within which to call bqsr from the input reads. All intervals will have a padding of 100 to get read records, and overlapping intervals will be combined. Interval files should be passed using the --interval-file option. This option can be used multiple times (e.g. "-L chr1 -L chr2:10000 -L chr3:20000+ -L chr4:10000-20000"). (default: None)
Pass supported bwa mem options as one string. The current original bwa mem supported options are -M, -Y and -T (e.g. --bwa-options="-M -Y") (default: None)
Suppress warning messages about system thread and memory usage. (default: None)
Don't generate SAM entries in the output if the entry's flag's meet this criteria. Criteria: (flag & filter != 0) (default: 0)
Filter SAM entries whose length of SA is not 0 (default: None)
Generate output BAM after bwa-mem. The output will not be co-ordinate sorted or duplicates will not be marked (default: None)
Do not perform the Mark Duplicates step. Return BAM after sorting. (default: None)
Add mate cigar (MC) and mate quality (MQ) tags to the output file. (default: None)
Assume the reads are sorted by queryname for Marking Duplicates. This will mark secondary, supplementary, and unmapped reads as duplicates as well. This flag will not impact variant calling while increasing processing times. (default: None)
Assume marking duplicates to be similar to Picard version 2.18.2. (default: None)
Monitor approximate CPU utilization and host memory usage during execution. (default: None)
The maximum offset between two duplicate clusters in order to consider them optical duplicates. Ignored if --out-duplicate-metrics is not passed. (default: None)
SM tag for read groups in this run. (default: None)
LB tag for read groups in this run. (default: None)
PL tag for read groups in this run. (default: None)
Prefix for the ID and PU tags for read groups in this run. This prefix will be used for all pairs of fastq files in this run. The ID and PU tags will consist of this prefix and an identifier, that will be unique for a pair of fastq files. (default: None)
Amount of padding (in base pairs) to add to each interval you are including. (default: None)
Run standalone BQSR. (default: None)
Flag alignments to strand 'f' or 'r' as failing quality-control (QC) with the failed QC flag 0x200. BS-Seq libraries are often to a single strand; other strands can be flagged as QC failures. Note: f == OT, r == OB. Valid options are 'f' or 'r' (default: None)
Turn off the default heuristic which marks alignments as failing QC if the longest match is less than 44% of the original sequence length. Alignments which fail this heuristic are also un-paired (default: None)
Performance Options:
- --bwa-nstreams BWA_NSTREAMS
- --bwa-cpu-thread-pool BWA_CPU_THREAD_POOL
- --gpuwrite
- --gpuwrite-deflate-algo GPUWRITE_DEFLATE_ALGO
- --gpusort
- --use-gds
- --memory-limit MEMORY_LIMIT
- --low-memory
Number of streams per GPU to use; note: more streams increases device memory usage (default: 4)
Number of threads to devote to CPU thread pool per GPU (default: 16)
Use one GPU to accelerate writing final BAM. (default: None)
Choose the nvCOMP DEFLATE algorithm to use with --gpuwrite. Note these options do not correspond to CPU DEFLATE options. Valid options are 0 and 3. Option 0 is faster while option 3 provides a better compression ratio. (default=0) (default: None)
Use GPUs to accelerate sorting and marking. (default: None)
Use GPUDirect Storage (GDS) to enable a direct data path for direct memory access (DMA) transfers between GPU memory and storage. Must be used concurrently with --gpuwrite. Please refer to Parabricks Documentation > Best Performance for information on how to set up and use GPUDirect Storage. (default: None)
System memory limit in GBs during sorting and postsorting. By default, the limit is half of the total system memory. (default: 62)
Use low memory mode; will lower the number of streams per GPU (default: None)
Common options:
- --logfile LOGFILE
- --tmp-dir TMP_DIR
- --with-petagene-dir WITH_PETAGENE_DIR
- --keep-tmp
- --no-seccomp-override
- --version
Path to the log file. If not specified, messages will only be written to the standard error output. (default: None)
Full path to the directory where temporary files will be stored.
Full path to the PetaGene installation directory. By default, this should have been installed at /opt/petagene. Use of this option also requires that the PetaLink library has been preloaded by setting the LD_PRELOAD environment variable. Optionally set the PETASUITE_REFPATH and PGCLOUD_CREDPATH environment variables that are used for data and credentials (default: None)
Do not delete the directory storing temporary files after completion.
Do not override seccomp options for docker (default: None).
View compatible software versions.
GPU options:
- --num-gpus NUM_GPUS
Number of GPUs to use for a run. GPUs 0..(NUM_GPUS-1) will be used.
The --in-fq option takes the names of two FASTQ files, optionally followed by a quoted read group. The FASTQ filenames must not start with a hyphen.
When using the --in-fq-list option a read group is required on each line of the input file.
Baseline bwa-meth: https://github.com/brentp/bwa-meth/
Bwa-meth manuscript: http://arxiv.org/abs/1401.1129