What's New?
Parabricks 4.0.0-1 is a major release with many significant changes. It streamlines how users get and set up the software and simplifies deployment on different platforms. If you are an existing Parabricks user, review this section to understand the major changes.
A license is no longer required to use Clara Parabricks. The container works out of the box once downloaded.
Clara Parabricks is free for
Academic research, or
Research by non-profit institutions, or
For development, test and evaluation purposes without use in production.
Users who would like to have Enterprise Support for Clara Parabricks can purchase NVIDIA AI Enterprise licenses, which provides full-stack support. To learn more about NVIDIA AI Enterprise, please visit https://www.nvidia.com/en-us/data-center/products/ai-enterprise/.
Parabricks 4.0.0-1 has a significantly reduced set of tools. We are focusing primarily on the following tools:
If you would like access to one or more of the tools that are no longer available, contact us in the developer forum.
Parabricks containers are compatible with WDL and NextFlow for building customized workflows, intertwining GPU- and CPU-powered tasks with different compute requirements, and deploying at scale.
These enable workflows to be deployed on cloud batch services as well as local clusters (e.g. SLURM) in a well managed process, pulling from a combination of Clara Parabricks and third-party containers and running these on pre-defined nodes.
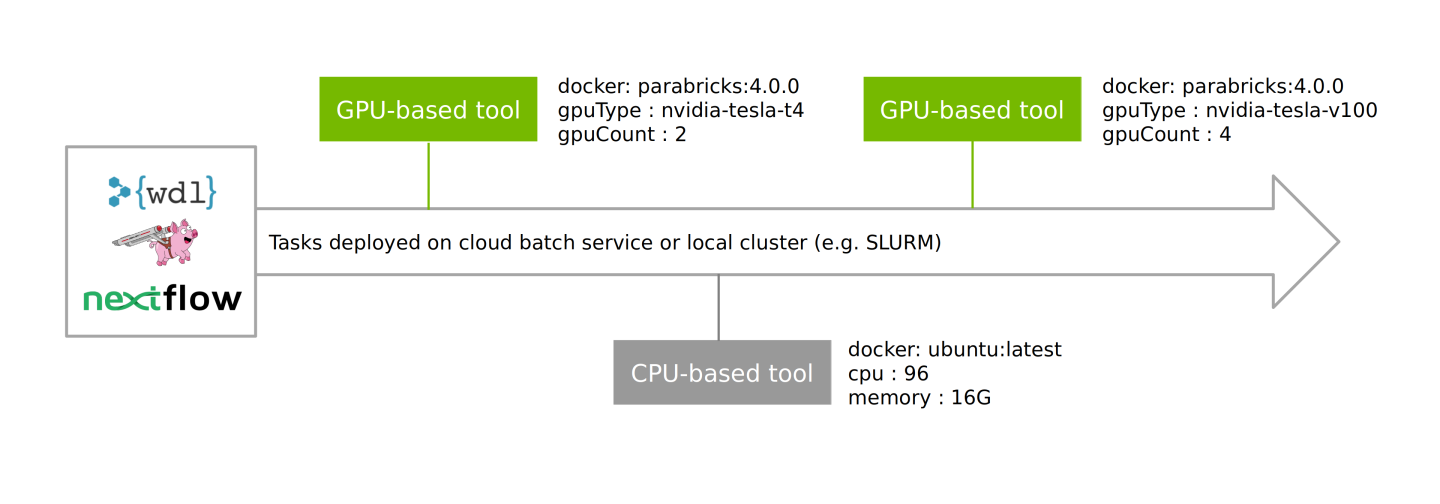
For further information on running these workflows, and to see the open-source reference workflows, which can be easily forked/edited, visit the Clara Parabricks Workflows repository. This repository includes recommended instance configurations for deploying the GPU-based tools on cloud and can be easily forked/edited for your own purposes.
deepvariant now implements DeepVariant v1.4.
haplotypecaller supports additional original HaplotypeCaller options.
deepvariant and deepvariant_germline support the
--channel-insert-sizeoption.starfusion now adds
PG:Z:MarkDuplicatesto each output BAM record.
Corrected rna_fq2bam sample code. The
--read-files-command zcat, while not a required parameter, is needed for correct operation with compressed FASTQ files.Updated the list of supported haplotypecaller options.
Corrected GATK sample code.
For further information see the Clara Parabricks datasheet.