Overview
Welcome to the trial of NVIDIA MegaMolBART on NVIDIA LaunchPad!
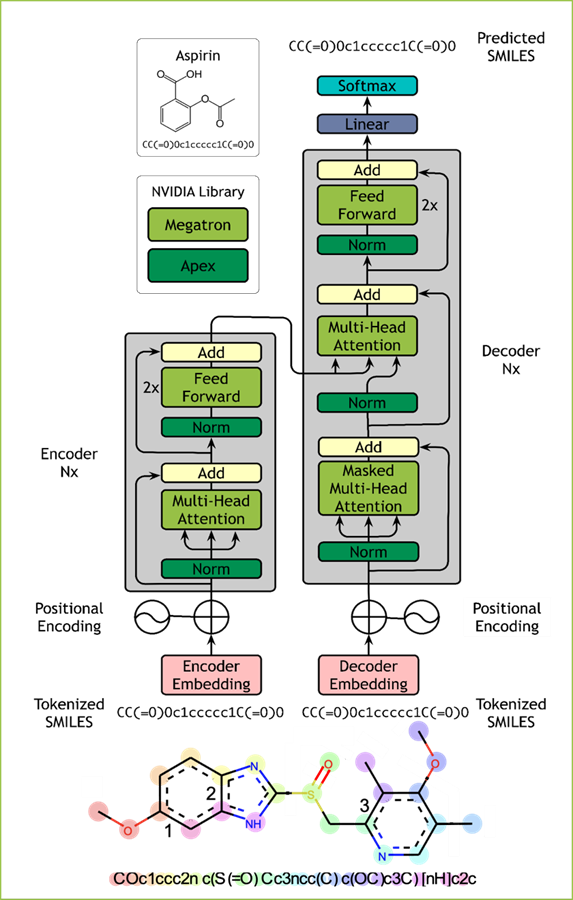
MegaMolBART is a deep learning model for small molecule drug discovery and cheminformatics based on SMILES. MegaMolBART uses NVIDIA’s Megatron framework, designed to develop large transformer models.
MegaMolBART relies on NeMo. NeMo provides a robust environment for developing and deploying deep learning models, including Megatron models. NeMo provides enhancements to PyTorch Lightning, such as hyperparameter configurabilityconfiguarbility with YAML files and checkpoint management. It also enables the development and training of large transformer models using NVIDIA’s Megatron framework, which makes multi-GPU, multi-node training with data parallelism, model parallelism, and mixed precision.
The ZINC-15 database is used for pre-training. Approximately 1.45 Billion molecules (SMILES strings) were selected from tranches meeting the following constraints: molecular weight <= 500 Daltons, LogP <= 5, reactivity level was “reactive”, and purchasability was “annotated”. SMILES formats, including chirality notations, are used as-is from ZINC.
During pre-processing, the compounds are filtered to ensure a maximum length of 512 characters. Train, validation, and test splits are randomly split using a seed as 99% / 0.5% / 0.5%. Data canonicalization and augmentation during training are performed using RDKIT via masking and SMILES randomization, as described previously.
The LaunchPad experience for BioNeMo_MegaMolBART will have two main objectives:
Showcase how one can perform model training using a chemical dataset
Configuring Weights and Biases for logging
Components/files for building and training the model
Launching the training job
A walkthrough of how to obtain learned embeddings for a set of molecules from MMB and make a downstream prediction model.
Generating new designs/molecules
Obtaining embedding and generating a prediction model
More information about MegaMolBART is available in the model guide.
The NeMo User Guide for more information on NeMo.