This Version of Parabricks is no longer available: This documentation page is for reference only. Versions 3.8 and earlier have been deprecated. We encourage you to use the latest version, for which no license is required. If you need access to this version of Parabricks to continue an ongoing project, please contact the Parabricks team at parabricks-support@nvidia.com.
- What's New?
- Getting Started with Clara Parabricks
- Software Overview
- Standalone Tools Overview
- Pipeline Tools Overview
- Tutorials
- How-Tos
- Technical Reference
How the Documentation is Organized
The documentation begins with this main page, which contains a brief introduction to Clara Parabricks: What it is, what it can do, and how it works.
The What's New? page covers what's changed since the previous release: new tools, improvements to existing tools, and bug fixes.
The Getting Started with Clara Parabricks page tells you everything you need to know to get started using Clara Parabricks:
What you need to use it
How to get it
How to install it
How to run it at peak performance
The Tutorials walk you through a single use of Clara Parabricks. First, you get some sample data so you can follow along and get the same output as shown. Then you convert a reference and FASTQ files to a BAM file, do variant calling on the BAM file, and produce a VCF file. Finally, you perform quality control on the VCF file and examine the resulting plots.
After an initial exposure in the tutorials, the How-Tos explore larger, more involved tasks, examining a wider variety of options, tools, and workflows. Owing to the larger data sets in use, a more capable hardware platform may be required (more GPUs, more memory, etc).
Finally, the Technical Reference contains reference documentation for each tool, organized both by category and alphabetically by tool name. You will learn how to compare the output of Clara Parabricks with the output from the baseline tools. You will also find a list of publications referencing Clara Parabricks, a list of frequently asked question, and pointers on getting more help and information.
What is Clara Parabricks?
Parabricks is a software suite for performing secondary analysis of next generation sequencing (NGS) DNA and RNA data. It delivers results at blazing fast speeds and low cost. Clara Parabricks can analyze 30x WGS (whole human genome) data in about 45 minutes, instead of 30 hours for other methods. Its output matches commonly used software, making it fairly simple to verify the accuracy of the output.
Why use Clara Parabricks?
Under the hood, Parabricks achieves this performance through tight integration with GPUs, which excel at performing data-parallel computation much more effectively than traditional CPU-based solutions. Parabricks was built from the ground up by GPU computing and Deep Learning experts who wanted to develop the fastest and most efficient possible implementation of common genomics algorithms used in secondary analysis.
Learn more at the Clara Parabricks developer page.
Software Overview
Parabricks is a software suite for genomic analysis. It delivers major improvements in throughput time for common analytical tasks in genomics, including germline and somatic analysis. The core of the Parabricks software is its data pipeline, which takes raw data and transforms it according to the user's requirements.
The Parabricks software supports the tools shown below:
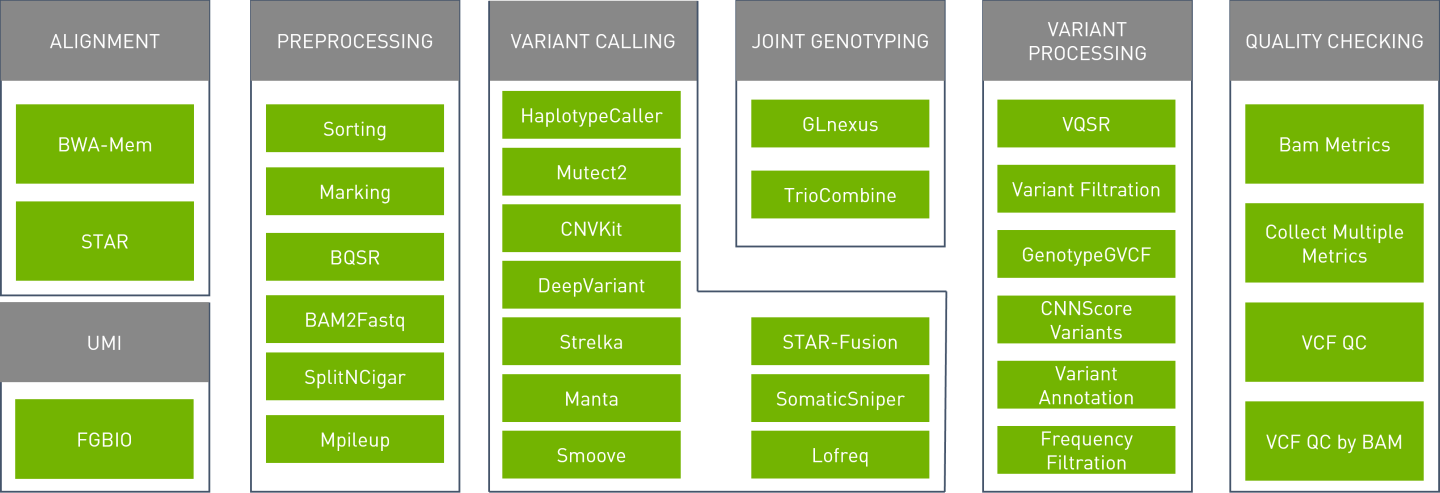
Parabricks Pipelines
Parabricks pipelines can simplify the steps required for processing. In Clara Parabricks, each pipeline is a collection of several individual tools that are commonly used together, all wrapped up as a single tool. For example, the deepvariant pipeline takes FASTA and FASTQ files as input and produces a VCF and BAM file as output. Internally, it runs BWA mem alignment, performs coordinate sorting, marks duplicates, and then runs DeepVariant.
Parabricks supports the pipelines shown below:
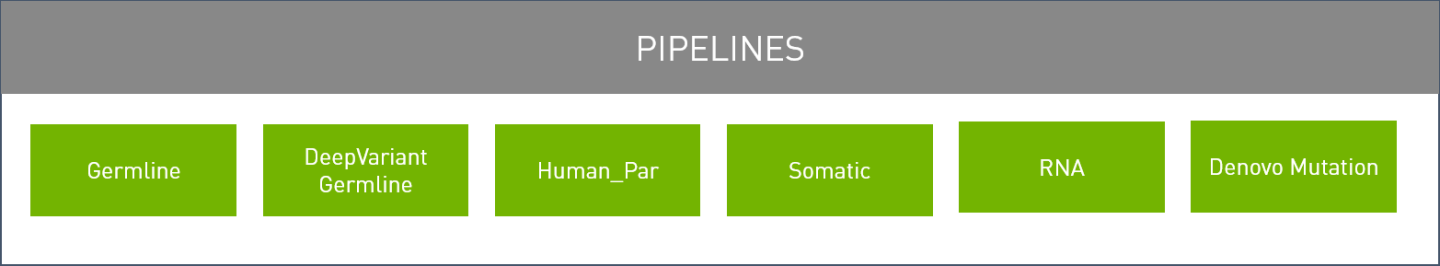

Parabricks can be configured to run specific accelerated tools or run full pipelines that are commonly used. The Software Tools section covers individual tools, and the Pipelines section covers commonly used pipelines.
NVIDIA Parabricks pipelines have been tested on Dell, HPE, IBM, and NVIDIA servers at Amazon Web Services, Google Cloud, Oracle Cloud Infrastructure, and Microsoft Azure.
How to Get Help
For technical support, updated user guides, and other Parabricks documentation, go to https://docs.nvidia.com/clara/#parabricks.
Answers to most FAQs can be found on the developer forum at https://forums.developer.nvidia.com/c/healthcare/Parabricks/290.
For further assistance, please contact us in one of the following ways:
Customers with PAID Parabricks licenses have direct access to support at https://nvid.nvidia.com or can contact EnterpriseSupport@nvidia.com
Users of FREE time-bound evaluation licenses can contact parabricks-eval-support@nvidia.com with troubleshooting questions.