1. Introduction
Clara Deploy SDK provides a container-based, cloud-native development and deployment framework for multi-AI, multi-domain workflows in smart hospitals–one platform managing and scaling Imaging, Genomics, and Video Processing workloads. It uses Kubernetes under the hood and enables developers and data scientists to define a multi-staged container-based pipeline.
The Clara Deploy SDK is a collection of containers that work together to provide an end to end medical image processing pipelines. The overall ecosystem can run on different cloud providers or local hardware with Pascal or newer GPUs.
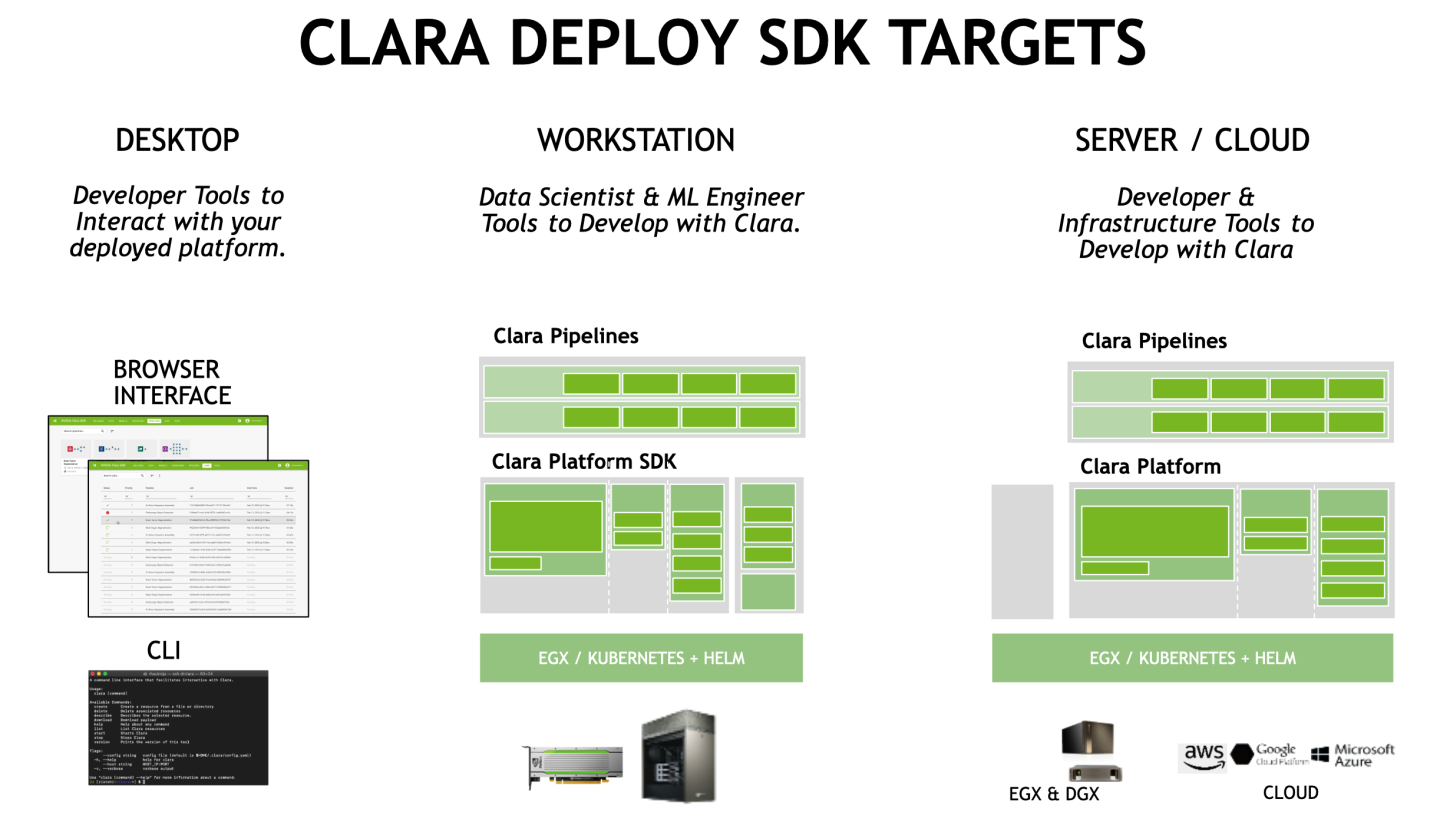
The Clara Deploy SDK is run via a collection of components that are deployed via Model Scripts/Helm Charts/Containers in NVIDIA GPU CLOUD (NGC) as pictured in the diagram below:
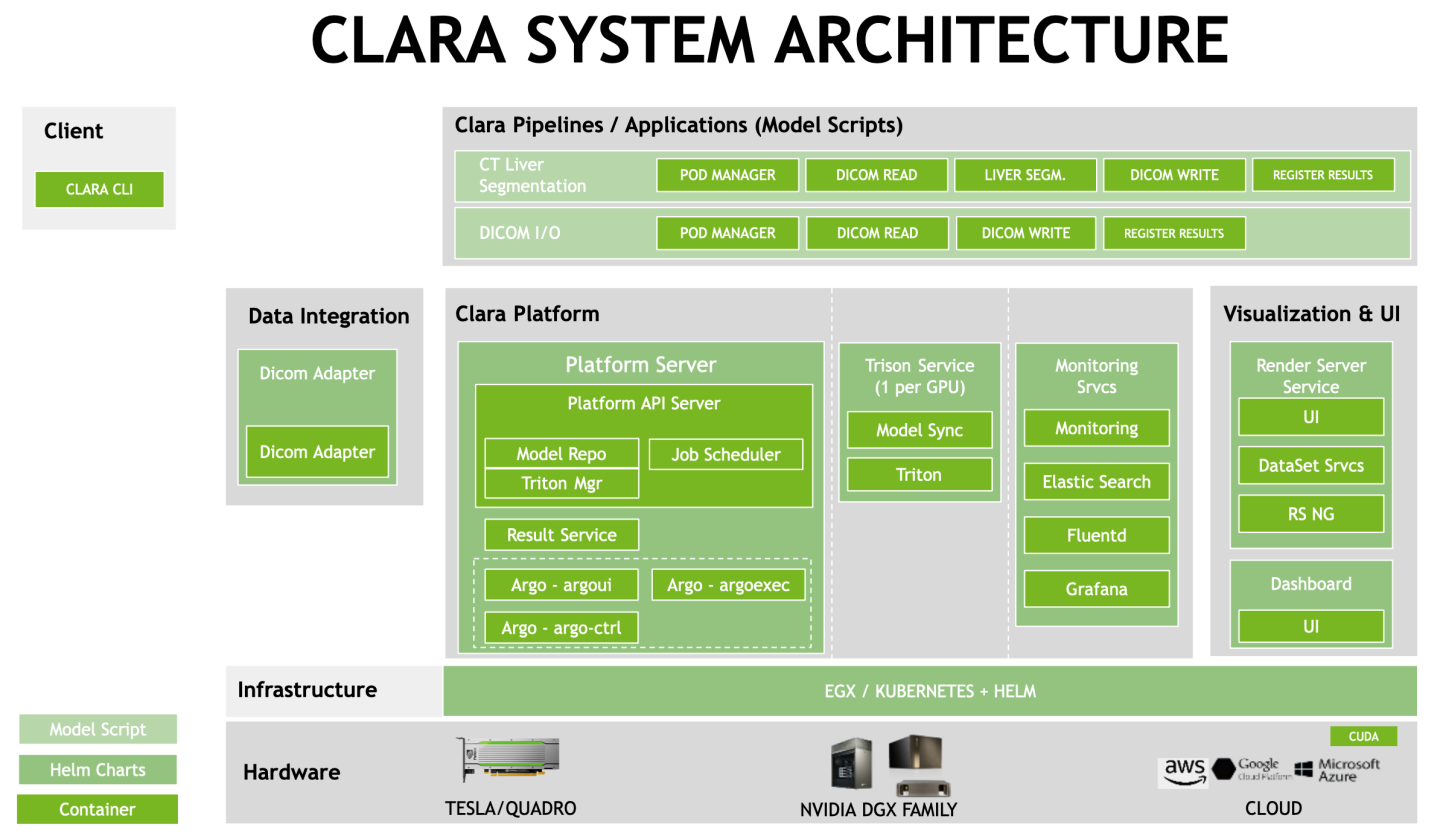
1.1.1.Clara Platform
Clara Platform component runs as the central part of the Clara Deploy SDK and controls all aspects of Clara payloads, pipelines, jobs, and results. Clara Platform acts as arbiter, router, and manager of data flowing in to and out of Clara Deploy SDK. Clara Platform is presented through Clara Platform API, enabling platform functionality to be consumed by external consumers (Clara CLI, UI, DICOM Adapter, external services) through Clara Platform API.
Clara Platform is responsible for the following:
Accepts and executes pipelines as jobs.
Deploys pipeline containers when needed via Helm, and removes them when those resources are needed elsewhere.
Being the source of truth for Clara system state.
1.1.1.1.Clara Platform API
Clara Platform provides a GRPC based interface enabling first- and third-party expansion of Clara Deploy SDK. Services such as DICOM Adapter, other PACS interfaces, or other satellite systems all interface with Clara Deploy SDK through the Clara Platform API. The API provides programmatic access to payload, pipeline, and job state; as well as the ability to create new pipelines, start pipeline jobs, and interface with the content of payloads.
As an example: Clara CLI, the provided utility for interfacing with Clara Deploy SDK from the command line, relies on Clara Platform API to provide the majority of its functionality. Similarly, DICOM Adapter relies on Clara Platform API to start, and monitor, pipeline jobs when DICOM data is sent to it.
1.1.2.DICOM Adapter
DICOM Adapter is the integration point between the hospital PACS and the Clara Deploy SDK. In the typical Clara Deploy SDK deployment, it is the first data interface to Clara. DICOM Adapter provides the ability to receive DICOM data, put the data in a payload, and trigger a pipeline. When the pipeline has produced a result, DICOM Adapter moves the result to a PACS.
1.1.3.Results Service
A Clara service that tracks all results generated by all pipelines. It bridges the results between the pipelines and the services delivering the results to external devices.
1.1.4.Clara Pipeline
A Clara pipeline is a collection of containers that are configured to work together to execute a medical image processing task. Clara publishes an API that enables any container to be added to a pipeline in the Clara Deploy SDK. These containers are Docker containers based on nvidia-docker with applications enhanced to support the Clara Container pipeline Driver.
1.1.5.Tensor RT Inference Server
The Tensor RT Inference Server is an inferencing solution optimized for NVIDIA GPUs. It provides an inference service via an HTTP or gRPC endpoint.
1.1.6.Render Server
Render Server provides visualization of medical data.
1.1.7.Clara I/O Model
The Clara I/O model is designed to follow standards that can be executed and scaled by Kubernetes. Payloads and results are separated to preserve payloads for restarting when unsuccessful or interrupted for priority, and to allow the inputs to be reused in multiple pipelines.
Payloads (inputs) are presented as read-only volumes.
Jobs are the execution of a pipeline on a payload.
Results (outputs, scratch space) are read-write volumes, paired with the jobs that created them in one-to-one relationships.
Datasets (reusable inputs) are read-only volumes.
Ports allow for application I/O.
Pipeline configurations are Helm charts.
Runtime container configuration is passed via a YAML file.