Mesh in ML Pipelines#
This chapter covers patterns for using PhysicsNeMo-Mesh in ML training pipelines.
Pipeline Positioning#
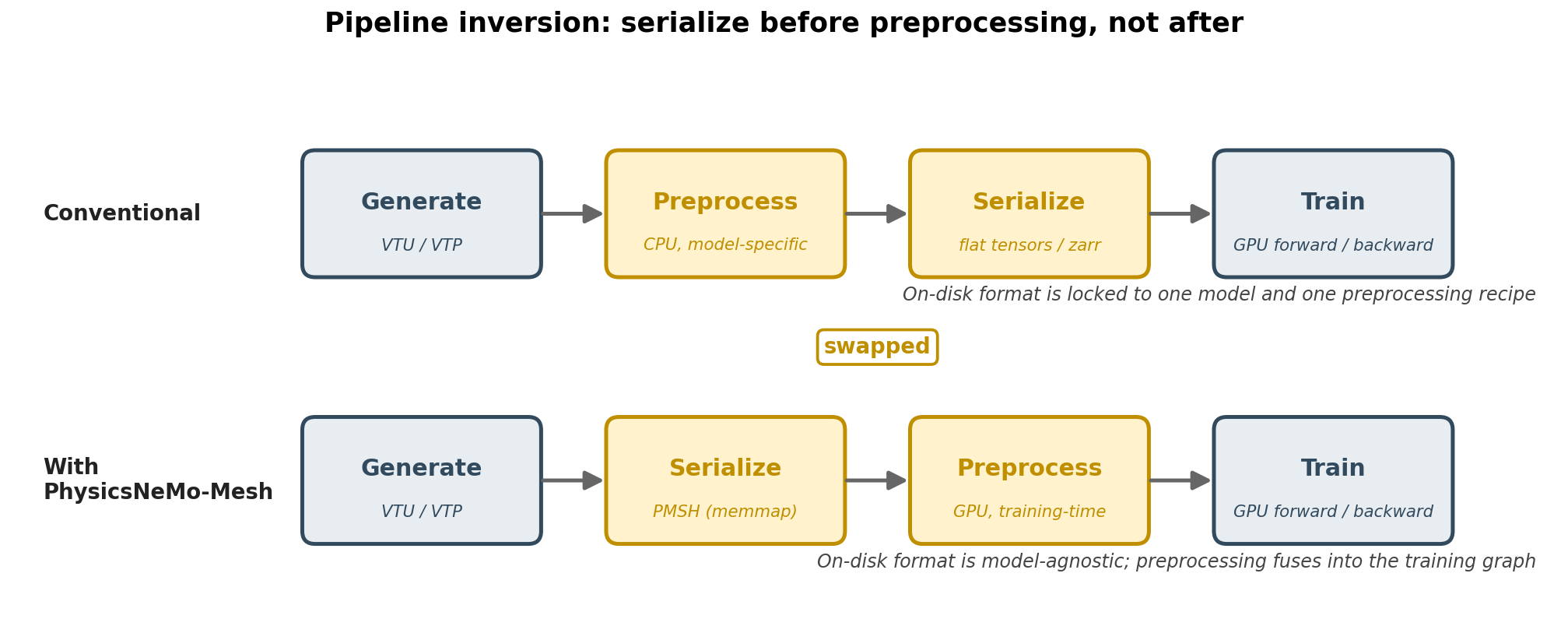
PhysicsNeMo-Mesh supports a serialize-then-preprocess ordering: convert
solver output to the model-agnostic *.pmsh format once, then
preprocess on-the-fly at training time on GPU. Because PMSH preserves
full mesh structure, the same serialized files can serve any downstream
model. Preprocessing is GPU-resident and autograd-compatible, so it can
run in parallel with the forward and backward pass and even fuse into a
single computational graph.
Training-Time Preprocessing#
Preprocessing fuses into the training step, not as a separate offline pass:
from physicsnemo.mesh import Mesh
def training_step(mesh: Mesh, model):
### Move to GPU and warm cached properties
mesh = mesh.to("cuda")
_ = mesh.cell_areas # warm cache before any compiled region
### Compute derived features at training time
mesh = mesh.compute_point_derivatives(keys="pressure", method="lsq")
### Forward pass: the model consumes the structured mesh
prediction = model(mesh)
loss = loss_fn(prediction, mesh.point_data["target"])
return loss
Standard PyTorch dataloader patterns (num_workers, pin_memory)
work unchanged. The next batch’s preprocessing and host-to-device
transfers run concurrently with the current batch’s forward and backward
pass.
Batching#
Different meshes typically have different numbers of points and cells, so they cannot be stacked into a single dense tensor without padding. Two common batching strategies:
Fixed-size padding. Pad every mesh in the batch up to a fixed size chosen to be larger than any expected mesh. Padding masks are stored as boolean tensors so the model can ignore the pad regions. Simplest when batch composition is known in advance.
Power-based padding. Pad up to the next power of two (or some other
discrete bucket size). Dramatically reduces the number of unique shape
signatures the model sees, which matters for torch.compile
shape-specialization caching: instead of recompiling for every distinct
batch size, the compiler only sees a small set of bucketed sizes. The
trade-off is some memory overhead for padding to the next bucket.
Both strategies are demonstrated in
tutorial 6.
The right choice depends on how heterogeneous the mesh sizes are in your
dataset and whether you are using torch.compile.
Feature Extraction#
Keeping mesh structure in the training loop enables computing new features from raw simulation data. A typical example: the DrivAerML dataset provides surface pressure and wall shear stress, but the classic boundary-layer-stability indicator is the streamwise pressure gradient \(dp/ds\), which is not directly available. With the mesh structure available at training time:
from physicsnemo.mesh.io import from_pyvista
import pyvista as pv
mesh = from_pyvista(pv.read("boundary_1.vtp")).to("cuda")
### Convert cell-centered fields to vertex-centered for differentiation
mesh = mesh.cell_data_to_point_data()
### Compute the surface pressure gradient (extrinsic, in 3D space)
mesh = mesh.compute_point_derivatives(
keys="pMeanTrim", gradient_type="extrinsic"
)
### Combine with the wall shear stress direction
grad_p = mesh.point_data["pMeanTrim_gradient"] # (n_points, 3)
wss = mesh.point_data["wallShearStressMeanTrim"] # (n_points, 3)
wss_hat = wss / wss.norm(dim=-1, keepdim=True).clamp(min=1e-8)
mesh.point_data["adverse_pg"] = (grad_p * wss_hat).sum(dim=-1)
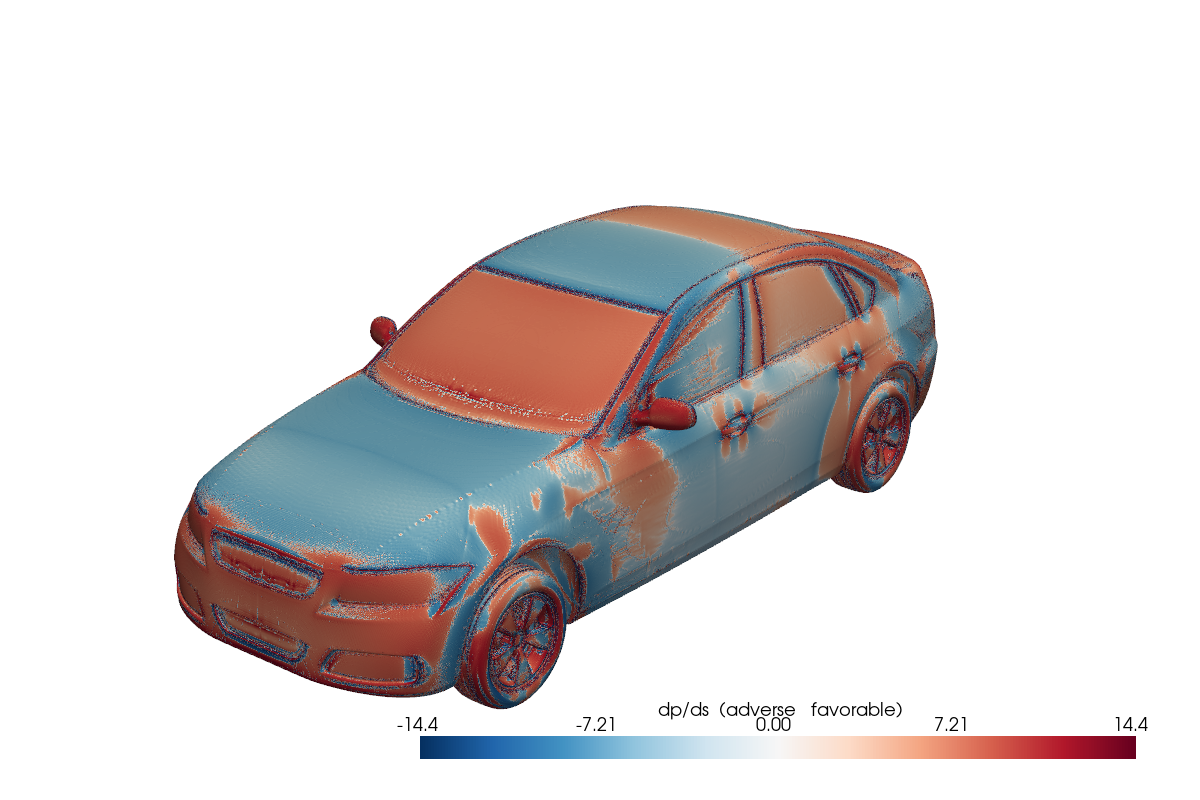

Fig. 19 Adverse pressure-gradient indicator on the DrivAerML car. Red regions are destabilizing; blue is favorable. Computed from raw simulation data at training time.#
The pattern is general: load raw data, convert between cell-centered and vertex-centered representations, compute gradients, combine with other fields. Each step is GPU-accelerated and autograd-compatible.
Data Augmentation#
For non-equivariant model architectures, augmenting the training data with random rotations, reflections, and translations of the same physical problem is a common way to improve generalization. With DomainMesh, this is a one-line operation that propagates consistently through the interior, every boundary, and the global metadata:
import math
import torch
dm = ... # DomainMesh loaded or constructed earlier
angle = torch.rand(1).item() * 2 * math.pi
dm_aug = dm.rotate(
angle=angle,
transform_point_data={"U": True},
transform_global_data={"U_inf": True},
)
The transform_*_data flags (covered in
Using Mesh) co-rotate vector and tensor fields
alongside the geometry. This is sometimes called quasi-equivariance:
the model is not structurally equivariant, but training on rotated
copies of each sample encourages it to behave approximately
equivariantly.
torch.compile compatibility#
The graph-breaking patterns and the three working patterns
(separate preprocessing from inner loops, pre-compute cached properties,
mode="reduce-overhead") are covered in
Using Mesh under torch.compile compatibility. They
apply equally to standalone preprocessing and to preprocessing fused
into a training step.
Future Directions#
PhysicsNeMo-Mesh is the first step in a data → layers → models progression toward physically-consistent, generalizable PDE surrogates:
Step 1 (data): PhysicsNeMo-Mesh and
DomainMeshprovide the semantics-rich data layer. This step is available today and works with any model architecture. For how to get your data intoMeshformat, see Creating and loading a Mesh.Step 2 (layers) (planned): TensorDict-compatible primitive layers in
physicsnemo.nn.layers(linear/MLP, attention, message-passing) that operate directly on nested semantic structure, without forcing an early flatten. An “off-ramp” collapses a TensorDict into a single concatenated tensor in a spatially-invariant manner so that non-equivariant downstream blocks can still consume the data when needed.Step 3 (models) (planned): New equivariant architectures built on top of data and layers.
Steps 2 and 3 are a roadmap; adopting Mesh and DomainMesh does not
require waiting for them. In the near term, several existing CAE datapipes
(DoMINO, Transolver, GeoTransolver, GLOBE) are being refactored to ingest and
produce Mesh TensorDicts as a drop-in replacement for CPU-bound PyVista/VTK
preprocessing.