Using Mesh#
Hands-on walkthroughs of the patterns below live at examples/minimal/mesh/ in the PhysicsNeMo repository (tutorials 1-6). For function- and class-level signatures, see the API reference.
Many of these operations run 10-100x faster on GPU than their CPU equivalents. This speedup makes it practical to run mesh preprocessing inside the training loop rather than as an offline step - and even to embed mesh operations within the model forward pass for end-to-end differentiability. For the full picture on training-time preprocessing, see Mesh in ML Pipelines.
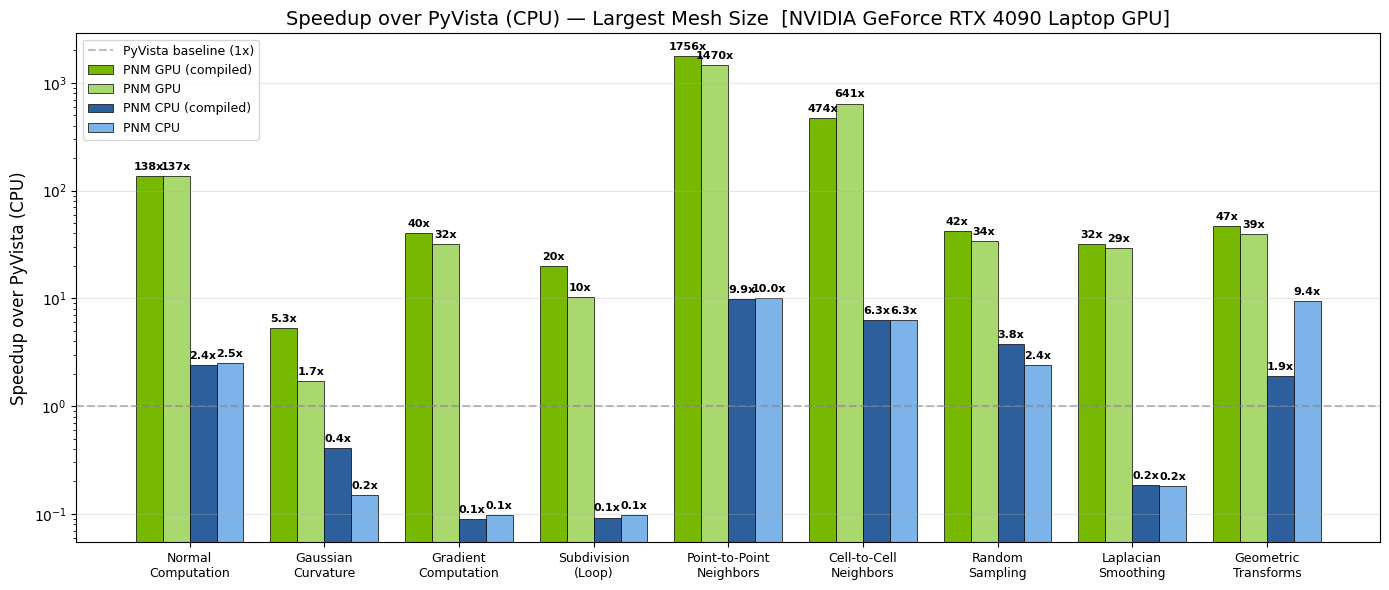
Fig. 14 Per-operation speedup of GPU PhysicsNeMo-Mesh (green) over CPU PyVista
(blue) on an NVIDIA GeForce RTX 4090 Laptop GPU. Darker shades show
torch.compile-d variants.#
Feature Overview#
The tables below summarize what is available, grouped by capability. Each group links to the corresponding API page.
Core data structure#
Capability |
Notes |
|---|---|
n-dimensional simplicial meshes |
Arbitrary manifold and spatial dimensions |
Point, cell, and global data containers |
Nested |
Memory-mapped serialization |
|
Mesh merging |
Concatenate multiple meshes |
Discrete calculus#
See physicsnemo.mesh.calculus.
Capability |
Notes |
|---|---|
Gradient, divergence |
LSQ (general-purpose) and DEC (rigorous discrete identities) |
Curl |
LSQ; 3D only |
Laplacian |
DEC, with cotangent weights |
Intrinsic vs extrinsic derivatives |
Tangent space vs ambient space, on embedded manifolds |
Geometry and curvature#
See physicsnemo.mesh.geometry and physicsnemo.mesh.curvature.
Capability |
Notes |
|---|---|
Cell areas, volumes, centroids |
Gram determinant method, dimensionally generic |
Cell and point normals |
Generalized cross product (cell); angle-area-weighted (point) |
Gaussian curvature |
Angle defect method |
Mean curvature |
Cotangent Laplacian |
Topology and adjacency#
See physicsnemo.mesh.boundaries and physicsnemo.mesh.neighbors.
Capability |
Notes |
|---|---|
Boundary and facet extraction |
Returns the boundary as a |
Watertight and manifold checks |
|
Adjacency: point-points, point-cells, cell-cells, cell-points |
Sparse (offsets, indices) encoding; |
Spatial queries and sampling#
See physicsnemo.mesh.spatial and physicsnemo.mesh.sampling.
Capability |
Notes |
|---|---|
BVH construction and traversal |
GPU-accelerated |
Point containment, nearest cell |
Per-query-point answers from a built BVH |
Barycentric interpolation |
Sample any field at arbitrary query points |
Random point sampling on cells |
Dirichlet-distributed barycentric coordinates within each cell |
Mesh surgery#
See physicsnemo.mesh.subdivision, physicsnemo.mesh.smoothing, physicsnemo.mesh.remeshing, and physicsnemo.mesh.repair.
Capability |
Notes |
|---|---|
Subdivision |
Linear (midpoint), Loop (C² smooth), Butterfly (interpolating) |
Laplacian smoothing |
Optional feature preservation |
Uniform remeshing |
Clustering-based; dimension-agnostic |
Cleanup ( |
Merges duplicate points, removes duplicate cells, drops unused vertices |
Quality metrics |
Aspect ratio, edge ratios, angles |
Transformations#
See physicsnemo.mesh.transformations.
Capability |
Notes |
|---|---|
Translate, rotate, scale |
Rotate: angle in 2D; angle-axis in 3D |
Arbitrary linear transform |
|
Projections#
See Transformations and Projections.
Capability |
Notes |
|---|---|
Project |
Drop spatial dimensions (e.g. 3D to 2D) |
Embed |
Add spatial dimensions (e.g. 2D to 3D) |
Extrude |
Increase manifold dimension by one (e.g. surface to volume) |
I/O and visualization#
See physicsnemo.mesh.io and physicsnemo.mesh.visualization.
Capability |
Notes |
|---|---|
PyVista interop |
|
Renderers |
Interactive PyVista (3D); matplotlib (2D / static 3D) |
Scalar coloring |
Auto L2-norm for vector fields |
Working with Meshes#
Creating meshes#
Four common ways to create a Mesh.
1. Directly from tensors. Useful for synthetic problems and tests:
import torch
from physicsnemo.mesh import Mesh
points = torch.tensor([
[0.0, 0.0],
[1.0, 0.0],
[1.0, 1.0],
[0.0, 1.0],
])
cells = torch.tensor([
[0, 1, 2],
[0, 2, 3],
])
mesh = Mesh(points=points, cells=cells)
2. From the built-in primitives library. Organized by category
(surfaces, volumes, planar, curves, procedural,
pyvista_datasets):
from physicsnemo.mesh.primitives.surfaces import sphere_icosahedral, torus
from physicsnemo.mesh.primitives.volumes import cube_volume
from physicsnemo.mesh.primitives.curves import helix_3d
sphere = sphere_icosahedral.load(radius=1.0, subdivisions=3)
donut = torus.load(major_radius=1.0, minor_radius=0.3)
cube = cube_volume.load(subdivisions=2)
helix = helix_3d.load(n_points=200, n_turns=4)
3. From a PyVista mesh. Handles every format PyVista supports (STL, VTK, VTU, VTP, PLY, OBJ, …) and triangulates non-simplicial cell types automatically:
import pyvista as pv
from physicsnemo.mesh.io import from_pyvista
pv_mesh = pv.read("path/to/geometry.stl")
mesh = from_pyvista(pv_mesh)
4. From a saved ``*.pmsh`` directory. See Serialization below.
Device management#
A single .to() call moves the entire mesh - geometry, every nested
data field, every cached property - to the target device or dtype:
mesh_gpu = mesh.to("cuda")
mesh_fp64 = mesh.to(torch.float64)
mesh_back = mesh_gpu.to("cpu")
mesh.pin_memory() is also available for fast asynchronous
host-to-device transfers in dataloaders.
Visualization#
mesh.draw() provides one-line visualization with automatic backend
selection: matplotlib for 2D, PyVista for 3D. Override with
backend="matplotlib" or backend="pyvista".
### Color by a point scalar field
mesh.draw(point_scalars="temperature", cmap="RdBu", show_edges=True)
### Color by a cell scalar field
mesh.draw(cell_scalars="pressure", cmap="viridis")
### Vector fields are auto-converted to L2-norm for coloring
mesh.draw(point_scalars="velocity", cmap="turbo")
In Jupyter notebooks, the PyVista backend supports interactive widgets via
pv.set_jupyter_backend("trame"). draw() returns the plotter / axes
object so you can customize before display.
Transformations#
Geometric transformations are pure functions: they return a new mesh and leave the original unchanged.
import numpy as np
mesh_translated = mesh.translate([1.0, 0.0, 0.0])
mesh_rotated = mesh.rotate(angle=np.pi / 4, axis=[0, 0, 1])
mesh_scaled = mesh.scale(2.0) # uniform
mesh_anisotropic = mesh.scale([2.0, 1.0, 0.5]) # per-axis
By default, transformations affect only the geometry (the points
tensor); attached field data is left unchanged. This is correct for
scalar fields like temperature (invariant under rotation) but wrong for
vector fields like velocity. Opt in to co-rotation via the
transform_*_data flags:
mesh_rotated = mesh.rotate(
angle=np.pi / 4,
axis=[0, 0, 1],
transform_point_data={"velocity": True},
transform_global_data={"U_inf": True},
)
Pass True to transform all compatible fields, or a dict /
TensorDict selecting specific fields. DomainMesh
uses the same flags to propagate transforms across an entire simulation
domain.
Note
In dimensions higher than 3, the angle-axis rotate formulation does
not generalize. Use mesh.transform(matrix) directly with an
arbitrary orthogonal matrix instead.
Topology and adjacency#
### Boundaries and facets
boundary = mesh.get_boundary_mesh() # the (n-1)-D outer boundary
facets = mesh.get_facet_mesh() # all (n-1)-D simplices, including interior
is_closed = mesh.is_watertight()
is_manifold = mesh.is_manifold()
### Adjacency queries (returned as sparse offset/index arrays)
point_neighbors = mesh.get_point_to_points_adjacency()
cell_neighbors = mesh.get_cell_to_cells_adjacency()
The adjacency objects use a sparse (offsets, indices) encoding.
Convert when you need a different representation:
neighbors_list = point_neighbors.to_list() # ragged list-of-lists
src, tgt = point_neighbors.expand_to_pairs() # (E,), (E,)
edge_index = torch.stack([src, tgt], dim=0) # PyG COO format: (2, E)
Discrete calculus: DEC vs LSQ#
Two complementary methods for discrete differential operators:
Discrete Exterior Calculus (DEC) is a mathematically rigorous framework where the discrete operators satisfy exact discrete versions of classical theorems (Stokes, Gauss-Bonnet, Helmholtz decomposition). It uses cotangent weights and circumcentric dual cells. See [Hirani2003] and [Desbrun2005].
Weighted least-squares (LSQ) is the practical CFD/FEM workhorse. Gradients are reconstructed by fitting a linear function to neighboring values; the result is exact for linear fields and first-order accurate in general.
Use DEC when you need exact discrete identities or are working with surface PDE solvers where the Laplace-Beltrami operator is the natural choice. Use LSQ for general-purpose CFD/FEM-style preprocessing where robustness on arbitrary topology matters more than exact discrete identities.
### Compute gradient of "pressure" via least-squares (default)
mesh = mesh.compute_point_derivatives(keys="pressure", method="lsq")
grad_p = mesh.point_data["pressure_gradient"] # shape (n_points, n_spatial_dims)
### Same with DEC (Discrete Exterior Calculus), for an intrinsic gradient
mesh = mesh.compute_point_derivatives(keys="pressure", method="dec")
For surfaces, you can choose between intrinsic (tangent-space) and extrinsic (ambient-space) derivatives:
mesh = mesh.compute_point_derivatives(keys="T", gradient_type="intrinsic")
mesh = mesh.compute_point_derivatives(keys="T", gradient_type="extrinsic")
For the underlying mathematics, see the physicsnemo.mesh.calculus API reference.
Differential geometry#
Curvature, normals, and tangent spaces are exposed as cached properties.
normals = mesh.point_normals # (n_points, n_spatial_dims)
K = mesh.gaussian_curvature_vertices # (n_points,)
H = mesh.mean_curvature_vertices # (n_points,)
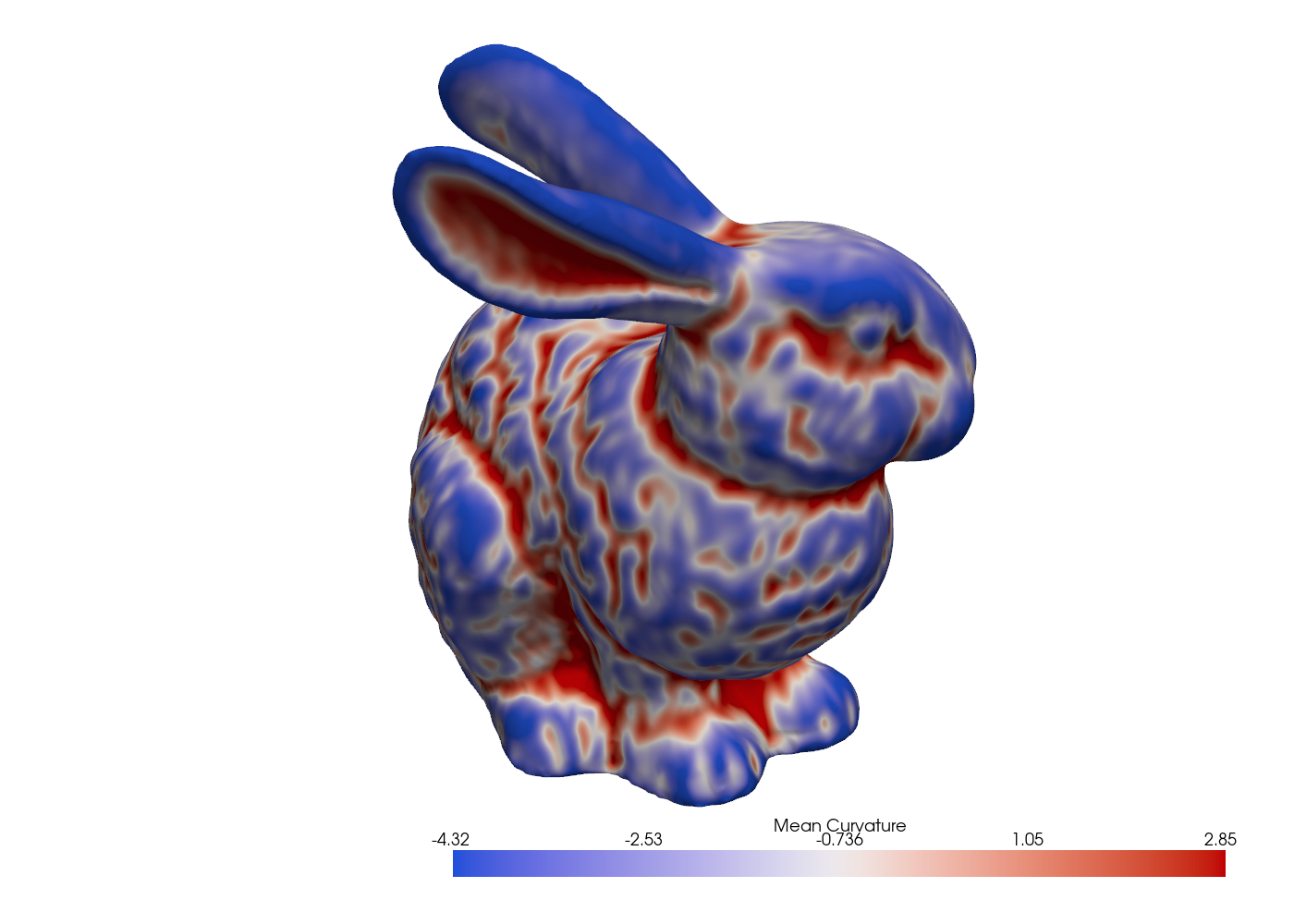
Fig. 15 Mean curvature on the Stanford bunny via the cotangent Laplacian.#
See the geometry and curvature API references for the full list of available properties and functions.
Spatial queries#
For point-in-mesh queries, nearest-cell searches, and barycentric interpolation, build a Bounding Volume Hierarchy once and reuse it for many queries:
from physicsnemo.mesh.spatial import BVH
from physicsnemo.mesh.sampling import find_containing_cells
bvh = BVH.from_mesh(mesh)
### BVH gives fast approximate candidates (AABB overlap)
query_points = torch.rand(1000, 3)
candidates = bvh.find_candidate_cells(query_points)
### Exact containment uses barycentric coordinate testing
cell_indices, bary_coords = find_containing_cells(mesh, query_points)
### Or sample any field at the query points (barycentric interpolation)
sampled = mesh.sample_data_at_points(query_points, data_source="points")
Mesh surgery#
Subdivision, smoothing, remeshing, and repair return new meshes:
### Subdivision
refined = mesh.subdivide(levels=2, filter="linear") # midpoint, no smoothing
smooth = mesh.subdivide(levels=2, filter="loop") # C² smooth
interp = mesh.subdivide(levels=2, filter="butterfly") # interpolating
### Smoothing
from physicsnemo.mesh.smoothing import smooth_laplacian
smoothed = smooth_laplacian(mesh, n_iter=10)
### Remeshing: rebuild from scratch with a target cell count
from physicsnemo.mesh.remeshing import remesh
coarse = remesh(mesh, n_clusters=5000)
### Repair: merge duplicate points, deduplicate cells, drop unused vertices
clean = mesh.clean()
For incremental quality fixes (instead of the all-in-one clean()), use
the targeted methods in
physicsnemo.mesh.repair.
Serialization#
By convention, Mesh objects are saved to *.pmsh directories and
DomainMesh objects to *.pdmsh directories. Unlike a single-file
format such as VTU, each save produces a directory containing one
memory-mapped tensor file per leaf field, with subfolders that mirror the
mesh’s data structure. This is convenient for data surgery: you can add,
drop, or replace individual fields directly on disk without reloading or
rewriting the entire mesh.
mesh.save("/path/to/mesh.pmsh")
mesh = Mesh.load("/path/to/mesh.pmsh", device="cuda")
Loading uses byte-for-byte memory mapping with near-zero parsing overhead.
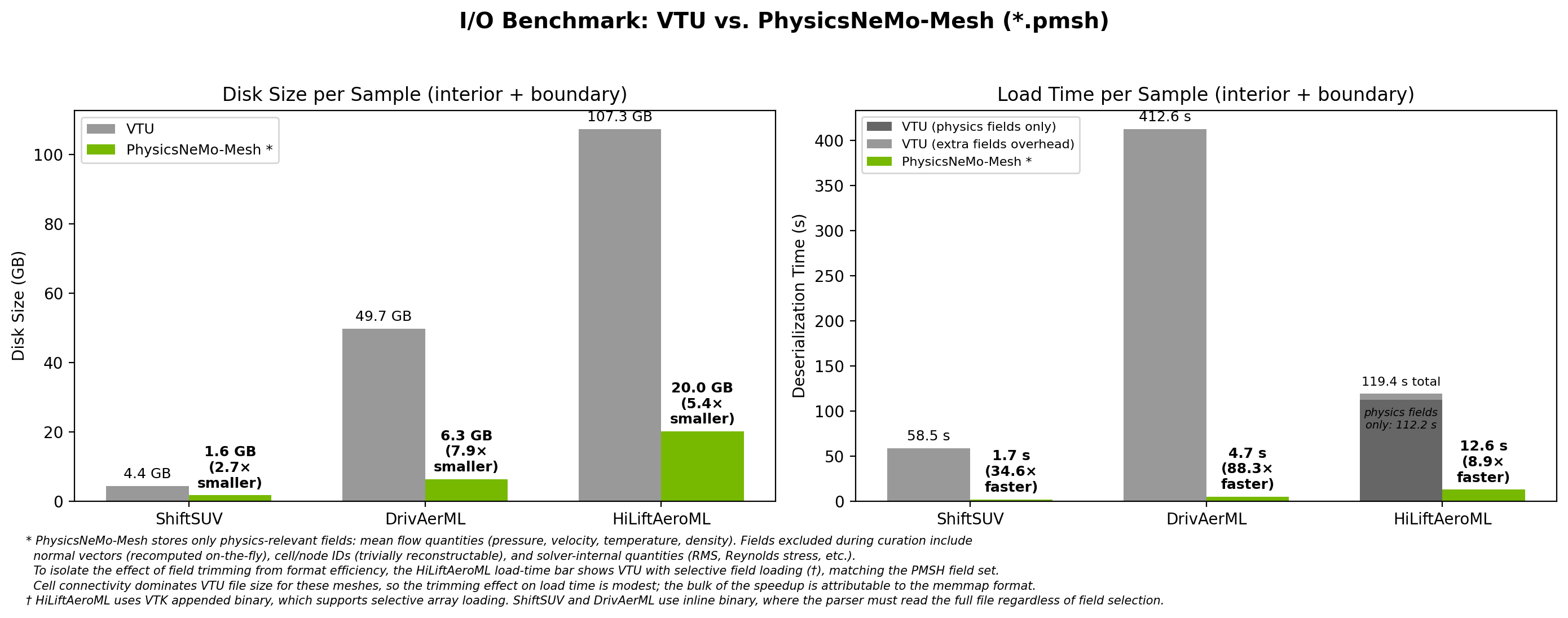
Fig. 16 PMSH is 9x to 88x faster to load than VTU and 3x to 8x smaller on disk across three production CFD datasets.#
The speedup comes from two sources: the user stores only the fields they
need (rather than the solver’s full output), and the memory-mapped format
avoids parsing overhead entirely. save() itself does not drop any
fields - it serializes the entire Mesh faithfully. The directory
layout also enables partial loading (read just the fields you need)
and on-disk surgery (swap a single field without rewriting the whole
mesh). See DomainMesh for the same pattern applied
to full simulation domains.
torch.compile compatibility#
Most operations are torch.compile-compatible, but some patterns cause
graph breaks:
scatter_add_operations (used in edge counting, facet extraction, adjacency).torch.wherewith variable-length output.torch.uniqueand similar operations with variable-sized outputs.
Three patterns work in practice:
1. Separate preprocessing from inner loops. Wrap mesh-topology computations in non-compiled code:
### Outside compiled region
neighbors = mesh.get_point_to_points_adjacency()
@torch.compile
def compute_laplacian(points, neighbor_indices, neighbor_offsets):
...
2. Pre-compute cached properties before entering compiled code. Cached
quantities like mesh.cell_areas are computed lazily on first access;
the first access inside a compiled function causes a graph break. Touch
them once outside first:
_ = mesh.cell_areas # warm cache
_ = mesh.cell_normals
@torch.compile
def my_loss(mesh):
...
3. Use ``mode=”reduce-overhead”`` when graph breaks are unavoidable. This minimizes per-graph-break dispatch cost.
Graph and Point Cloud Conversion#
For the conceptual relationship between meshes, graphs, and point clouds - including why a geometrically embedded graph carries more information than an abstract one - see Graphs, Point Clouds, and Geometric Embedding in the Concepts chapter.
For a higher-dimensional mesh, two natural graph representations are available:
The edge graph (the 1-skeleton) treats mesh vertices as nodes and mesh edges as edges. Best when data lives on vertices, as in most graph-neural-network surrogates.
The dual graph treats cell centroids as nodes, with edges between cells that share a facet. Best when data lives on cells, as in finite-volume CFD where fluxes are exchanged at cell-cell interfaces.
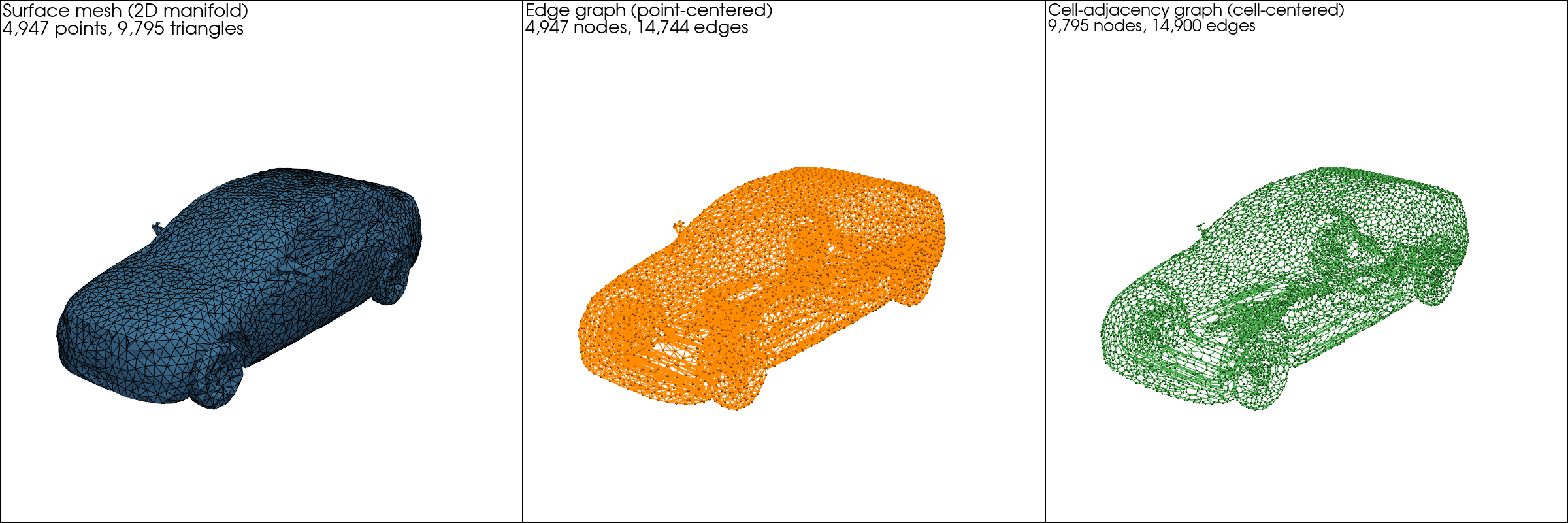

Fig. 17 The DrivAerML car: surface mesh (left), edge graph (center,
vertex-centered), and dual graph (right, cell-centered). All three are
first-class Mesh objects.#
surface = ... # Mesh[2, 3]: triangles in 3D
edge_graph = surface.to_edge_graph() # Mesh[1, 3]: vertex-centered
dual_graph = surface.to_dual_graph() # Mesh[1, 3]: cell-centered
point_cloud = surface.to_point_cloud() # Mesh[0, 3]: no connectivity
Each converted result is itself a Mesh and can use any of the
operations covered earlier. These conversion methods are documented in the
Mesh API reference.
PyTorch Geometric interop#
For workflows already built around PyG, adjacency objects convert to
edge_index format:
adjacency = mesh.get_point_to_points_adjacency()
src, tgt = adjacency.expand_to_pairs()
edge_index = torch.stack([src, tgt], dim=0) # shape (2, n_edges)
from torch_geometric.nn import GCNConv
x = mesh.point_data["features"]
y = GCNConv(in_channels=x.shape[-1], out_channels=64)(x, edge_index)